This is a sequel to the previous post, https://karencfisher.com/2021/09/24/recognizing-covid-19-in-chest-x-rays-using-a-convolutional-neural-network/
In the previous post, we implemented a relatively simple convolutional neural network to classify chest x-rays as to whether showing COVID-19 infection, contrasting with normal cases. While that model seemed successful, with high accuracy, sensitivity, specificity, and AUC scores (all greater than 90%), looking at class activation maps it was not clear that the model was really specific as to the lung region. In many cases in fact it seemed to be looking anywhere else in the images.
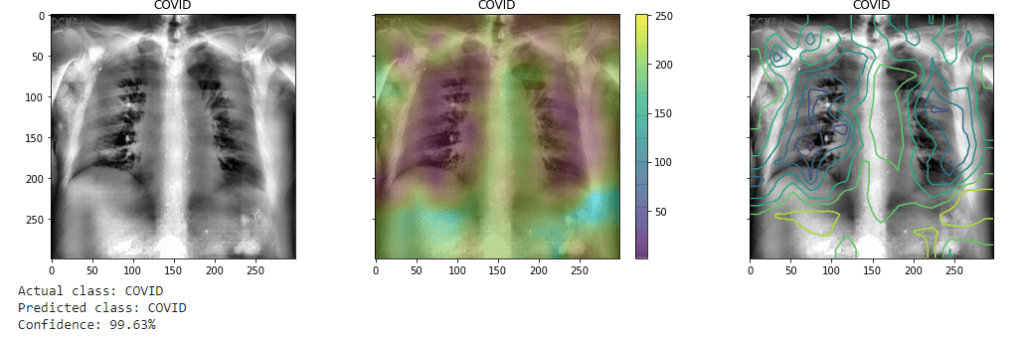
(As before the CAM is visualized as both heatmap and contour plots superimposed over the image.) It may also give the greatest weights to even annotations on the images.
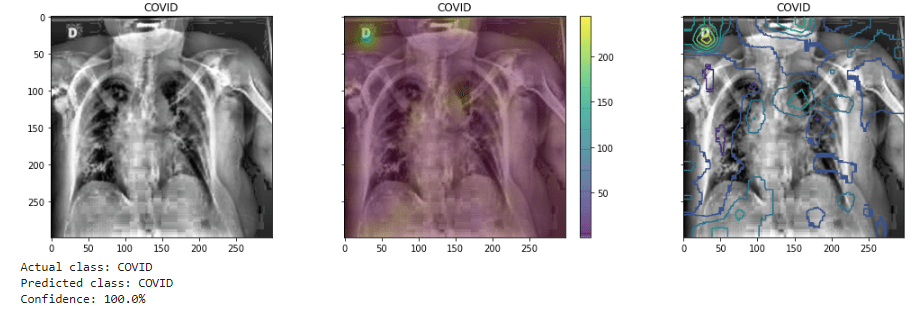
This has been noted by other researchers: for example “The convenience of working with an open dataset often comes with a price—the image quality could vary among samples in the collection and noisy or irrelevant annotations may be included in some images.” [1] The model’s attention wanders, as it were, finding variations other than we might want it to look at and weighing them heavier in classifying the image.
One solution that has been suggested has been to essentially mask off the less relevant regions of the image.[2] One may compare the approach perhaps with training a dog to play with a ball — and not other objects which may be nearby. One simply removes or masks off distractions.
The enhanced model presented now will propose a more complex model. Specifically two models: a segmentation model (U-Net architecture) to predict a mask for each image, the output of which is superimposed over the original images and fed through a deeper classifier (Densenet121) utilizing transfer learning. Using a pre-trained model such as Densenet allows greater accuracy with less available training data (as is often the case in medical datasets.

Lung Masking and Segmentation With U-Net
In order to mask out areas other than lungs in the x-rays, a segmentation model serves as the first layer. Image segmentation essentially classifies each pixel as being of one another class. It could be of a specific object — perhaps an automobile, a person, or animal — as opposed to the background of the image. Semantic segmentations highlights objects of a type in an image. In this case, the model classifies each pixel in the x-ray image as either part of lung, or not.

The resulting from the classification is a segmentation map. Each pixel is classified as 1 (the white region in a monochrome image) for lung, or 0 (the black region) for background or anything else. This segmentation map may then be used to essentially mask out anything in the image that is not lung simply by multiplying it with the original image pixel by pixel. This results in retaining the pixel values of the image in the selected area, while the rest are zeroed out.

A variation on an autoencoder network, the UNet architecture was developed specifically for use in biomedical image segmentation. In an autoencoder network, an image is first progressively down sampled, to a very concentrated latent representation, and then progressively up sampled to the original dimensions. This can produce a more abstracted or more generalized representation of the original image (other applications of autoencoders include denoising images as it loses some detail). In it the UNet version there are “skip connections” between the encoding and decoding layers of the same levels of down and up sampling. As a result it silhouettes, one might say, the specific detected object in the image. Utilizing an architecture as illustrated below, the model is trained on a dataset of lung segmentation which includes 700 chest X-ray images and corresponding segmentation maps as ‘ground truth.’ [3] The resultant model had an accuracy (dice accuracy) of roughly 96%. (Dice accuracy is a form of “intersection over union”: calculating the ratio between the number of pixels in both the predicted and truth-ground maps.)
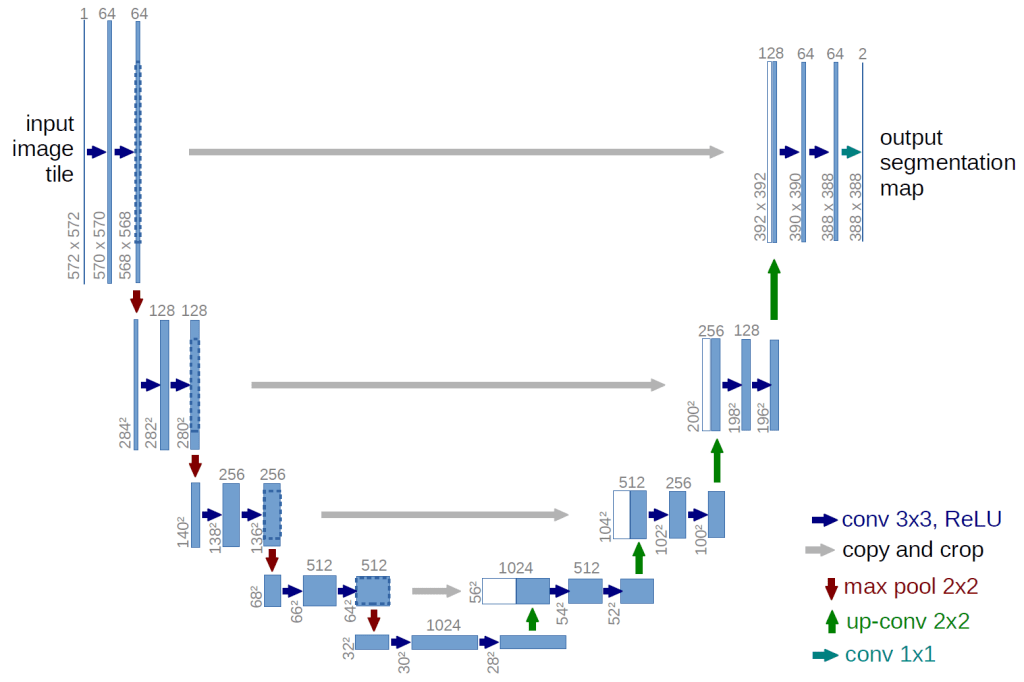
Transfer Learning with Densenet121
One of the more difficult of developing effective machine learning models for medical application is often the relative lack of available data. Highly performing deep learning models in computer vision require large volumes of images for training. However, often times datasets for medical applications are often, relatively speaking, small — in the order of perhaps around 30,000 images to be divided into training, validation, and testing datasets. By comparison state of the art image classification models, such as Resnet, Inception net, or Densenet, are typically trained on datasets such as imagenet, which contains over a million images for training and testing sorted into a thousand classes.
One approach to this is to utilize what is known as “transfer learning.” This consists of leveraging an advanced, state of the art, image classifier (such as Densenet), trained on vast numbers of images, for a more specialized purpose. The earlier stages of a convolutional neural network essentially extracts low level features, such as edges, which are common to all images. Hence, what those networks have “learned” is applicable to even types of images for which they were not necessarily explicitly trained: both cats and lungs have edges, for example. So that extensive training can be applied to a new domain, or a “downstream task.”
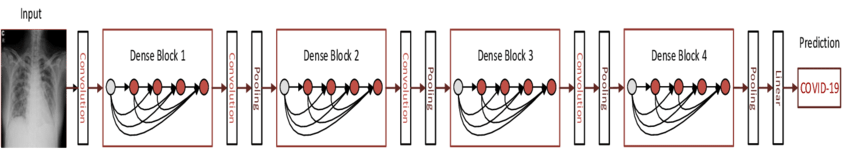
The model consists of a series of dense blocks, which are layers of convolutional layers densely connected to one another, each layer taking as input all the previous layers in the block. It then feeds the resultant feature maps to a fully connected layer (or set of layers) which perform classification (essentially logistic regression), producing a probability distribution of which class the image falls into.
For transfer learning, the fully connected layer (or layers) is replaced with ones which are for the specific application. All or at least the earlier convolutional layers are “frozen,” and only the upper and/or newly added layers are trained on the specific task. In this case, the upper most dense block is also “fine tuned,” as well as the final layers are trained on the X-ray images.
Model Evaluation
The final performance of the model on the validation and test datasets have been approximately 96%. More specifically, sensitivity (the rate of true positive predictions) has shown to be 93%, and specificity (the rate of true negative predictions) about 97%. In other words, it does somewhat better on predicting negative cases (the majority class in the datasets). However, this is overall a good outcome as the dataset is fairly imbalanced (1:3 ratio of positive vs. negative examples).

Additionally an alternative dataset, the QaTa-COVID19 dataset, was used for further conformation of the model. [6, 7] In that test, the accuracy overall was 93.5%, with a sensitivity of 95% and specificity of 93%.
In this final figure there are examples of predictions by the model on both COVID and non-COVID examples, with there respective probabilities. Overlaying with the images I have also superimposed saliency maps (shown both as heatmap and contour plots) to attempt to highlight the areas the model is now giving weight to those classifications. It is (as expected) now focused on the lungs area, and perhaps may indicate in positive cases the focus of damage from the virus. While it seems still inconclusive, such could be of use in medical diagnosis and also prognosis into determining the extent of damage and form that possibly the likely progression of the illness.

References
Project Github repository: https://github.com/karencfisher/COVID19
[1] Chun-Fu Yeh, Hsien-Tzu Cheng, Andy Wei, Hsin-Ming Chen, Po-Chen Kuo, Keng-Chi Liu, Mong-Chi Ko, Ray-Jade Chen, Po-Chang Lee, Jen-Hsiang Chuang, Chi-Mai Chen, Yi-Chang Chen, Wen-Jeng Lee, Ning Chien, Jo-Yu Chen, Yu-Sen Huang, Yu-Chien Chang, Yu-Cheng Huang, Nai-Kuan Chou, Kuan-Hua Chao, Yi-Chin Tu, Yeun-Chung Chang, Tyng-Luh Liu. A Cascaded Learning Strategy for Robust COVID-19 Pneumonia Chest X-Ray Screening. Page 3. https://arxiv.org/pdf/2004.12786.pdf
[2] Ibid., p. 3
[3] The dataset is compiled by the National Library of Medicine, National Institutes of Health, Bethesda, MD, USA and Shenzhen No.3 People’s Hospital, Guangdong Medical College, Shenzhen, China. See also Candemir S, Jaeger S, Palaniappan K, Musco JP, Singh RK, Xue Z, Karargyris A, Antani S, Thoma G, McDonald CJ. Lung segmentation in chest radiographs using anatomical atlases with nonrigid registration. IEEE Trans Med Imaging. I have obtained the dataset from https://www.kaggle.com/nikhilpandey360/chest-xray-masks-and-labels
[4] Olaf Ronneberger, Philipp Fischer, and Thomas Brox. U-Net: Convolutional Networks for Biomedical Image Segmentation. https://arxiv.org/pdf/1505.04597.pdf
[5] Kumar, Saurabh & Mishra, Shweta & Singh, Sunil. (2020). Deep Transfer Learning-based COVID-19 prediction using Chest X-rays. EURASIP Journal on Advances in Signal Processing volume 2021. https://asp-eurasipjournals.springeropen.com/articles/10.1186/s13634-021-00755-1
[6] A. Degerli, M. Ahishali, M. Yamac, S. Kiranyaz, M. E. H. Chowdhury, K. Hameed, T. Hamid, R. Mazhar, and M. Gabbouj, Covid-19 infection map generation and detection from chest X-ray images. Health Inf Sci Syst 9, 15 (2021). https://doi.org/10.1007/s13755-021-00146-8 I obtained this dataset from https://www.kaggle.com/aysendegerli/qatacov19-dataset
[7] A. Degerli, M. Ahishali, S. Kiranyaz, M. E. H. Chowdhury, and M. Gabbouj, Reliable Covid-19 detection using chest x-ray images. 2021 IEEE International Conference on Image Processing (ICIP), 2021, pp. 185-189, https://doi.org/10.1109/ICIP42928.2021.9506442.